Nature发文报道丨世界上首次实现跨膜孔蛋白的精确从头设计
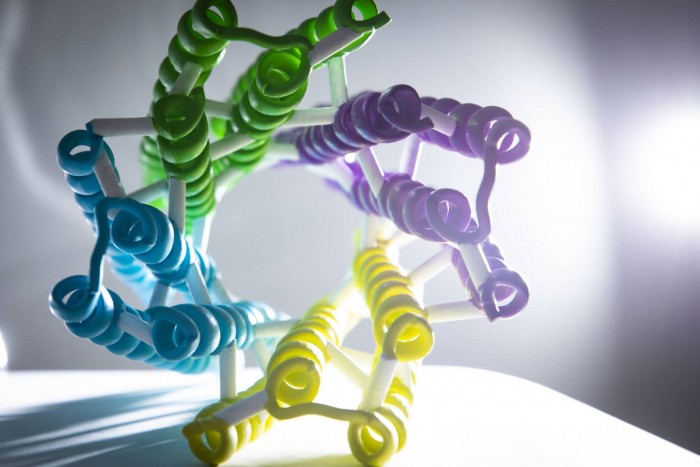

北京时间8月26日23时,Nature杂志在线发表西湖大学生命科学学院卢培龙研究员课题组与华盛顿大学David Baker等课题组合作的人工设计跨膜蛋白质的最新研究:《跨膜孔蛋白的计算机辅助设计》(Computational Design of Transmembrane Pores)。该研究在世界上首次实现了跨膜孔蛋白的精确从头设计。
华盛顿大学徐纯福博士和西湖大学生命科学学院卢培龙研究员为该文的共同第一作者, 卢培龙研究员、华盛顿大学William A. Catterall教授和David...
卢培龙博士入选《麻省理工科技评论》“35岁以下35人”中国榜单

卢培龙
年龄:31 岁
职位:西湖大学生命科学学院PI
获奖理由:他首次实现了多次跨膜蛋白三维结构的精确设计。
蛋白质设计是结构生物学的重要分支和新兴的前沿学科,需要生物物理学、生物化学、合成生物学以及计算生物学等多学科的交叉融合。
卢培龙通过计算生物学手段模拟蛋白质极性残基在膜环境内部形成的相互作用,设计了能够在膜环境中稳定存在的蛋白质三维结构,并进行重组表达、生化性质测定、三维结构解析等一系列实验来进行验证。
卢培龙利用这一方法成功设计了多种具有极高热稳定性...
Keep Your Eyes on the Prize: A Life Connected with Science -------- By Ronald D. Vale
"I feel that my greatest prize has been the privilege of a life connected with science. I have enjoyed innumerable discoveries, both my own and those of others, and I have met many kind and interesting people along the way. My best advice to young scientists is to keep your eyes on these pri...
Programmable design of orthogonal protein heterodimers
Protein–protein interaction specificity can be achieved using extensive and modular side-chain hydrogen-bond networks. We used the Crick generating equations to produce millions of four-helix backbones with varying degrees of supercoiling around a central axis, identified thos...
Prof. David Baker shared his insights on de novo protein design with PNAS, talking about transmembrane protein design.
"If you want to build an airplane, you don’t start by modifying a bird; instead, you understand the first principles of aerodynamics and build flying machines from those principles. As we get closer to solving the protein folding problem, we can now design completely new proteins from ...
新华社报道:科学家用计算机设计制造出跨膜蛋白质
新华社华盛顿3月2日电(记者周舟)
美国华盛顿大学一个科研团队借助计算机设计,制造出了跨膜蛋白质,它可像天然膜蛋白那样形成多蛋白复合体,可应用在疫苗设计等领域。
膜蛋白是生物膜中所含的蛋白质,嵌在细胞和细胞器的膜中。有些膜蛋白的两端暴露于膜的内外表面,被称作跨膜蛋白质。它们可作为物质跨膜转运的通道,许多药物设计将其作为靶标。
本次研究论文发表在最新一期《科学》杂志上。论文第一作者、华盛顿大学贝克实验室的卢培龙对新华社记者说,他们使用一种叫做“罗塞塔”的计算机程序,设计了能在膜环境...
The coming of age of de novo protein design

There are 20200 possible amino-acid sequences for a 200-residue protein, of which the natural evolutionary process has sampled only an infinitesimal subset. De novo protein design explores the full sequence space, guided by the physical principles that underlie prote...
De novo Protein Design---Science's Breakthrough of the Year, 2016
"Designing new proteins from scratch has been a hit-or-miss activity. It’s easy enough to write any desired DNA code, but researchers have had no way of knowing how the novel strings of amino acids encoded by this DNA would fold into complex 3D shapes. That’s a problem, beca...
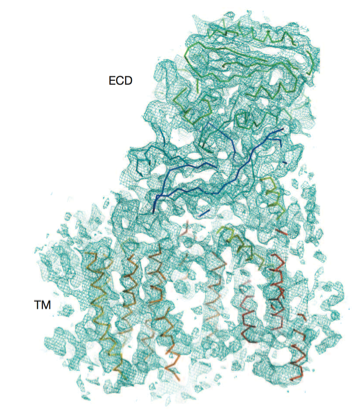
Three-dimensional structure of human γ-secretase
The γ-secretase complex, comprising presenilin 1 (PS1), PEN-2, APH-1 and nicastrin, is a membrane-embedded protease that controls a number of important cellular functions through substrate cleavage. Aberrant cleavage of the amyloid precursor protein (APP) results in aggregation of amyl...