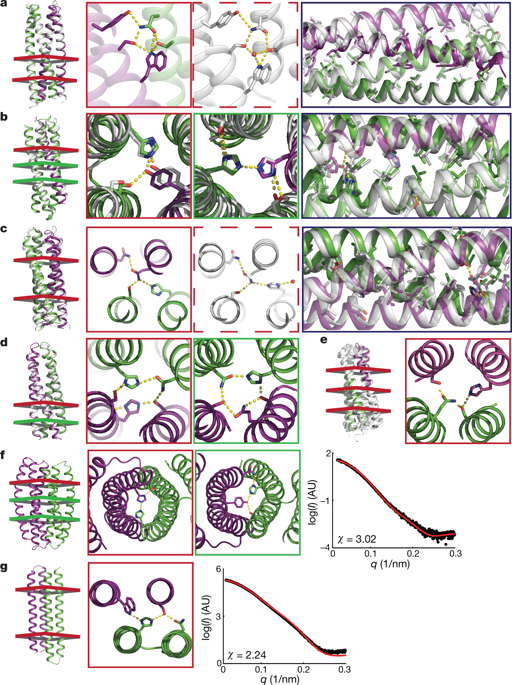
Protein–protein interaction specificity can be achieved using extensive and modular side-chain hydrogen-bond networks. We used the Crick generating equations to produce millions of four-helix backbones with varying degrees of supercoiling around a central axis, identified those accommodating extensive hydrogen-bond networks, and used Rosetta to connect pairs of helices with short loops and to optimize the remainder of the sequence. Of 97 such designs expressed in Escherichia coli, 65 formed constitutive heterodimers, and the crystal structures of four designs were in close agreement with the computational models and confirmed the designed hydrogen-bond networks. In cells, six heterodimers were fully orthogonal, and in vitro—following mixing of 32 chains from 16 heterodimer designs, denaturation in 5 M guanidine hydrochloride and reannealing—almost all of the interactions observed by native mass spectrometry were between the designed cognate pairs. The ability to design orthogonal protein heterodimers should enable sophisticated protein-based control logic for synthetic biology, and illustrates that nature has not fully explored the possibilities for programmable biomolecular interaction modalities.
https://www.nature.com/articles/s41586-018-0802-y