"Designing new proteins from scratch has been a hit-or-miss activity. It’s easy enough to write any desired DNA code, but researchers have had no way of knowing how the novel strings of amino acids encoded by this DNA would fold into complex 3D shapes. That’s a problem, because for proteins, shape dictates function. Recently, however, computational biologists have made heady progress in designing computer programs that accurately predict how designer proteins will fold. Those advances made possible this year’s surge in designer proteins."
http://www.sciencemag.org/news/2016/07/protein-designer-aims-revolutionize-medicines-and-materials
创建: Dec 10, 2018 | 13:21
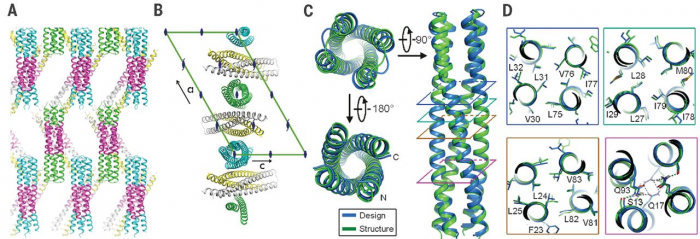
The computational design of transmembrane proteins with more than one membrane-spanning region remains a major challenge. We report the design of transmembrane monomers, homodimers, trimers, and tetramers with 76 to 215 residue subunits containing two to four membrane-spanning regions and up to 860 total residues that adopt the target oligomerization state in detergent solution. The designed proteins localize to the plasma membrane in bacteria and in mammalian cells, and magnetic tweezer unfolding experiments in the membrane indicate that they are very stable. Crystal structures of the designed dimer and tetramer-a rocket-shaped structure with a wide cytoplasmic base that funnels into eight transmembrane helices-are very close to the design models. Our results pave the way for the design of multispan membrane proteins with new functions.
http://science.sciencemag.org/content/359/6379/1042.long
创建: Dec 10, 2018 | 13:43
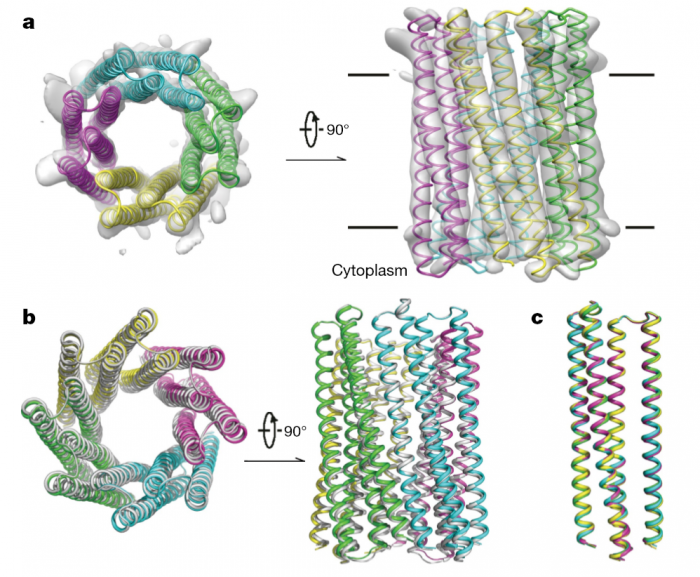
Transmembrane channels and pores have key roles in fundamental biological processes1 and in biotechnological applications such as DNA nanopore sequencing2–4, resulting in considerable interest in the design of pore-containing proteins. Synthetic amphiphilic peptides have been found to form ion channels5,6, and there have been recent advances in de novo membrane protein design7 ,8 and in redesigning naturally occurring channel-containing proteins9,10. However, the de novo design of stable, well-defined transmembrane protein pores that are capable of conducting ions selectively or are large enough to enable the passage of small-molecule fluorophores remains an outstanding challenge11,12. Here we report the computational design of protein pores formed by two concentric rings of α-helices that are stable and monodisperse in both their water-soluble and their transmembrane forms. Crystal structures of the water-soluble forms of a 12-helical pore and a 16-helical pore closely match the computational design models. Patch-clamp electrophysiology experiments show that, when expressed in insect cells, the transmembrane form of the 12-helix pore enables the passage of ions across the membrane with high selectivity for potassium over sodium; ion passage is blocked by specific chemical modification at the pore entrance. When incorporated into liposomes using in vitro protein synthesis, the transmembrane form of the 16-helix pore—but not the 12-helix pore—enables the passage of biotinylated Alexa Fluor 488. A cryo-electron microscopy structure of the 16-helix transmembrane pore closely matches the design model. The ability to produce structurally and functionally well-defined transmembrane pores opens the door to the creation of designer channels and pores for a wide variety of applications.
创建: Sep 27, 2020 | 17:15
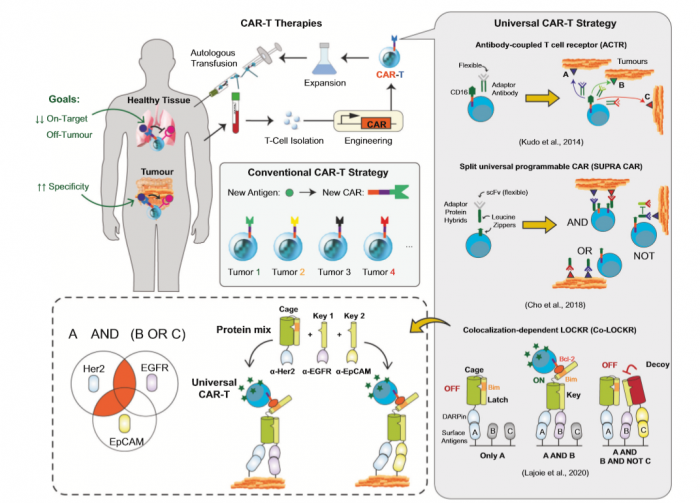
Protein logic gates acting at posttranscriptional levels are unique in being amenable to the control of both intracellular and intercellular biological processes. In a recent paper published in Science, Lajoie et al. engineered highly versatile colocalization-dependent protein switch that are activated upon detection of logic combinations (e.g., AND, OR and NOT) of specific cell surface antigens, providing another important asset for “universal” CAR-T therapies.
创建: Sep 27, 2020 | 17:26